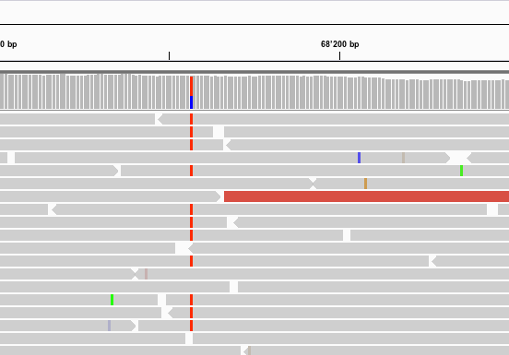
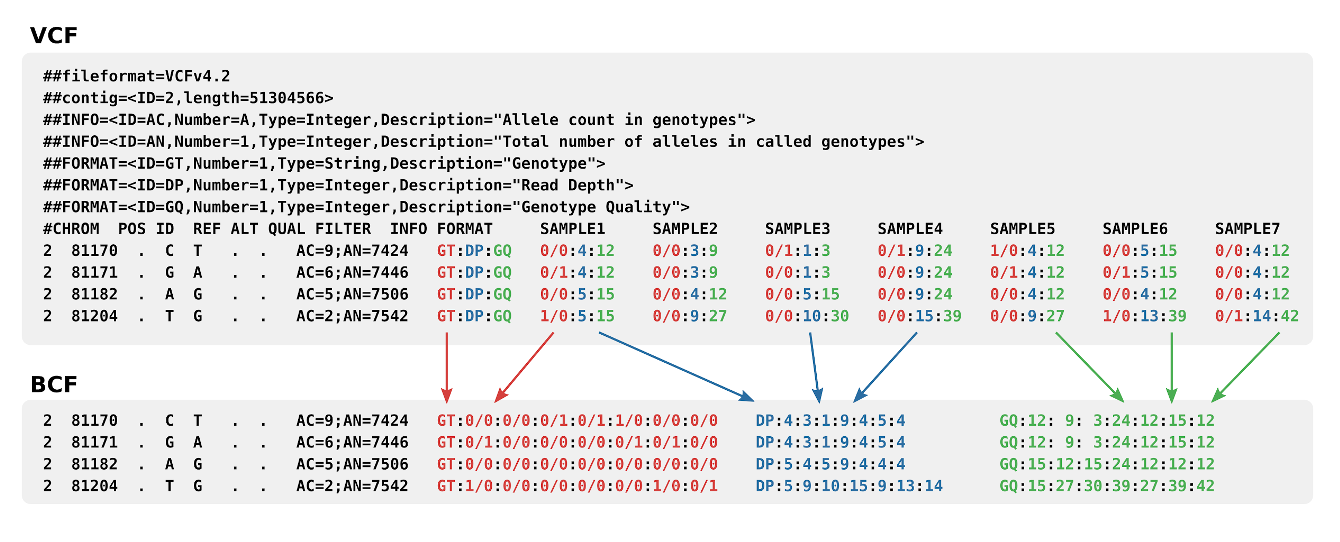
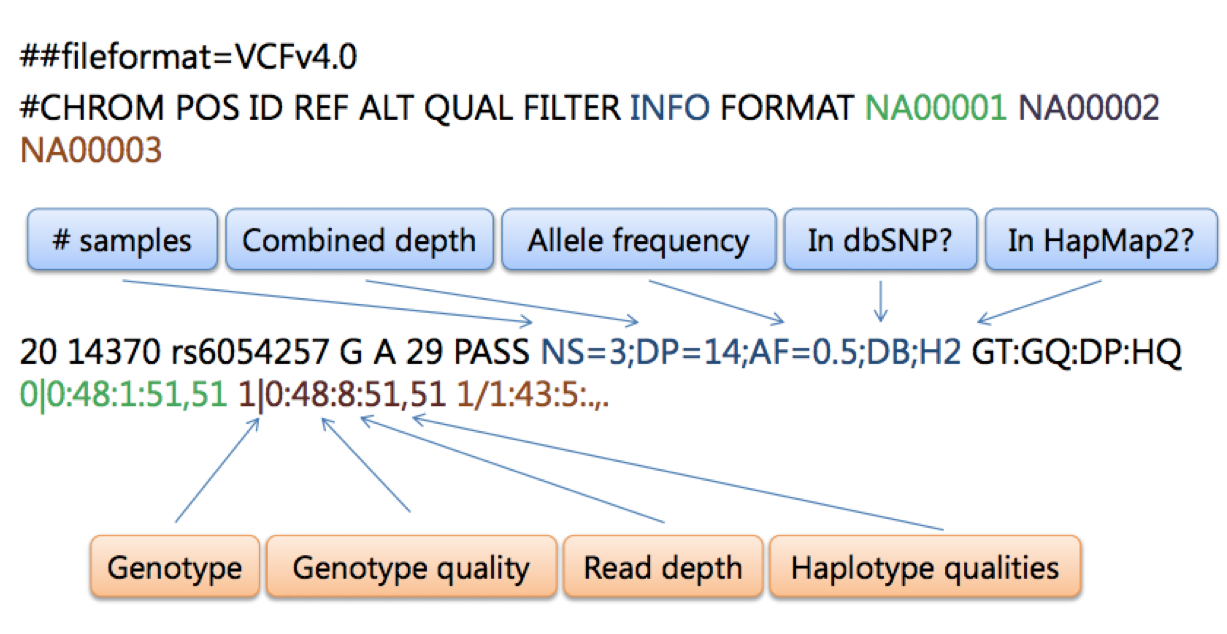
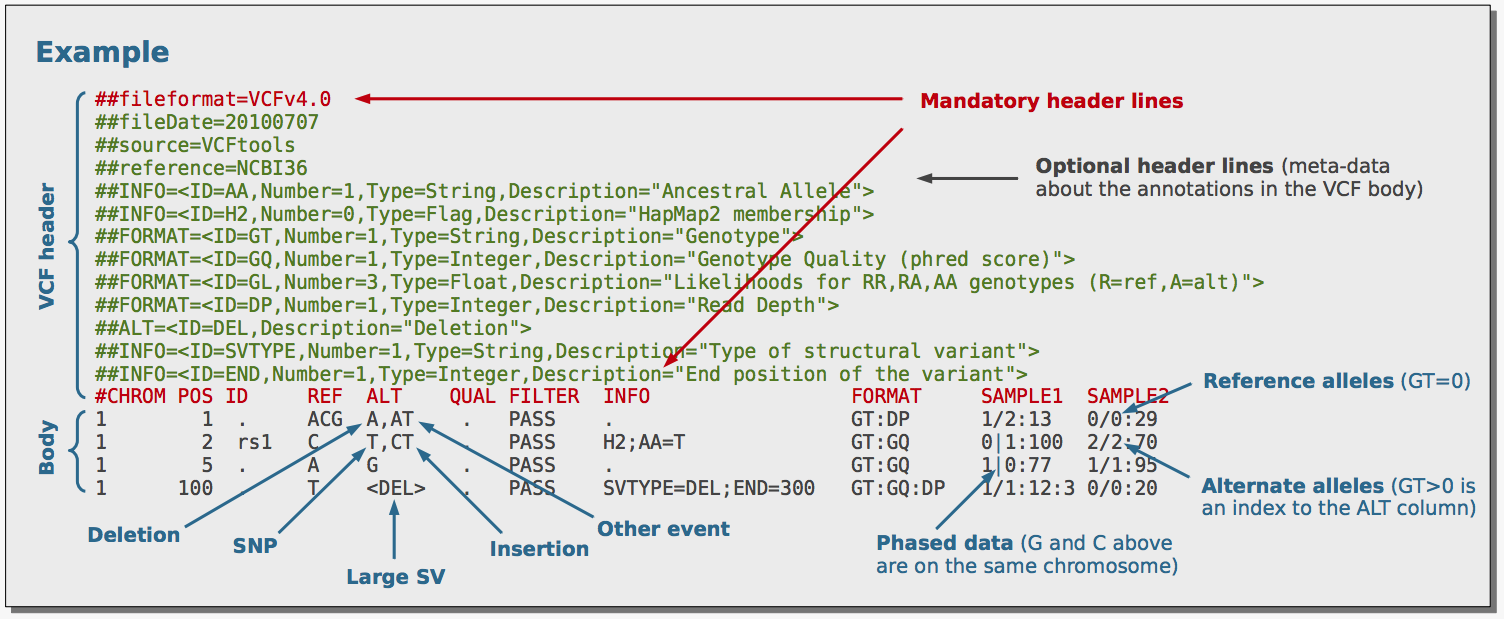
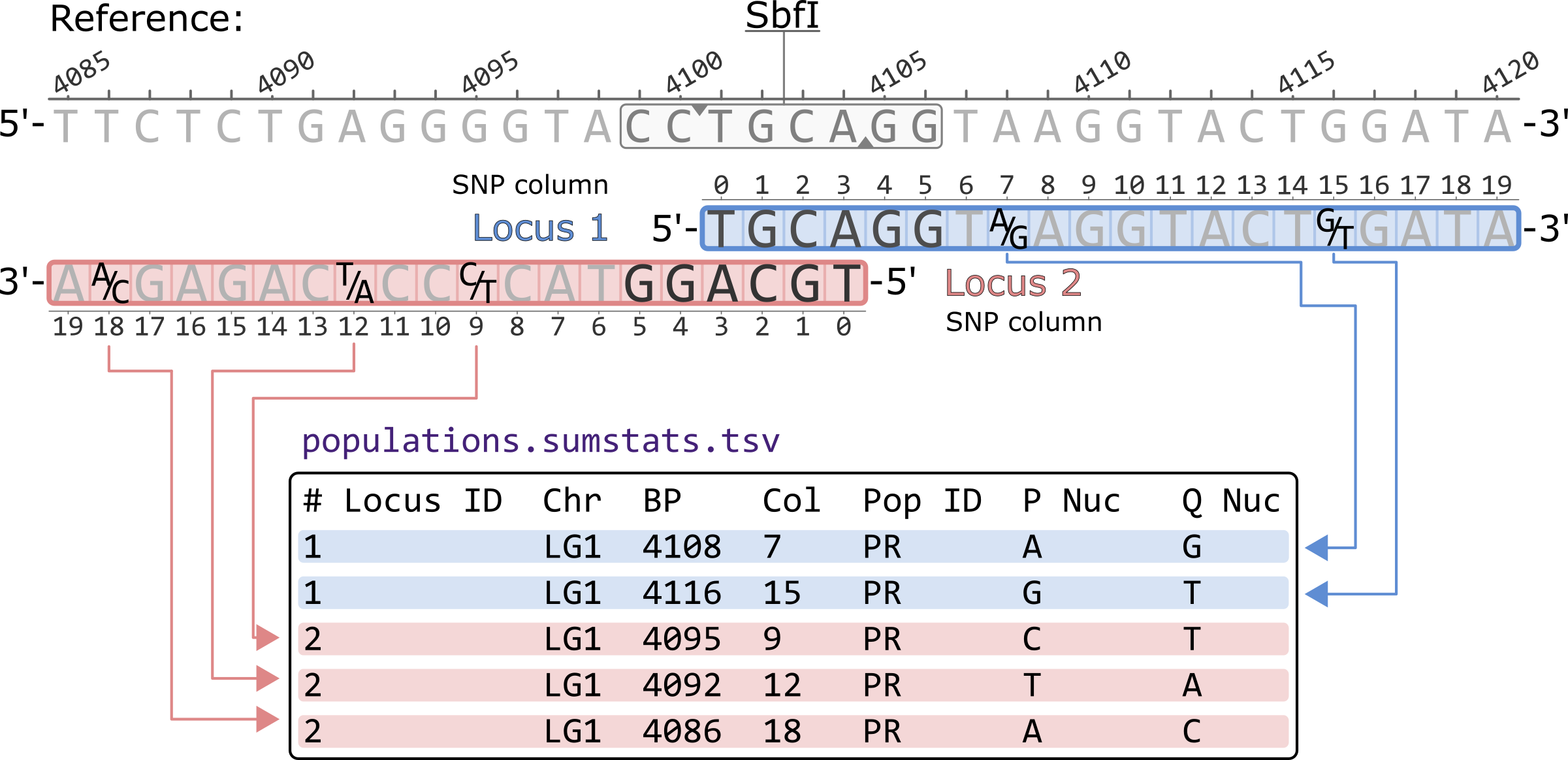
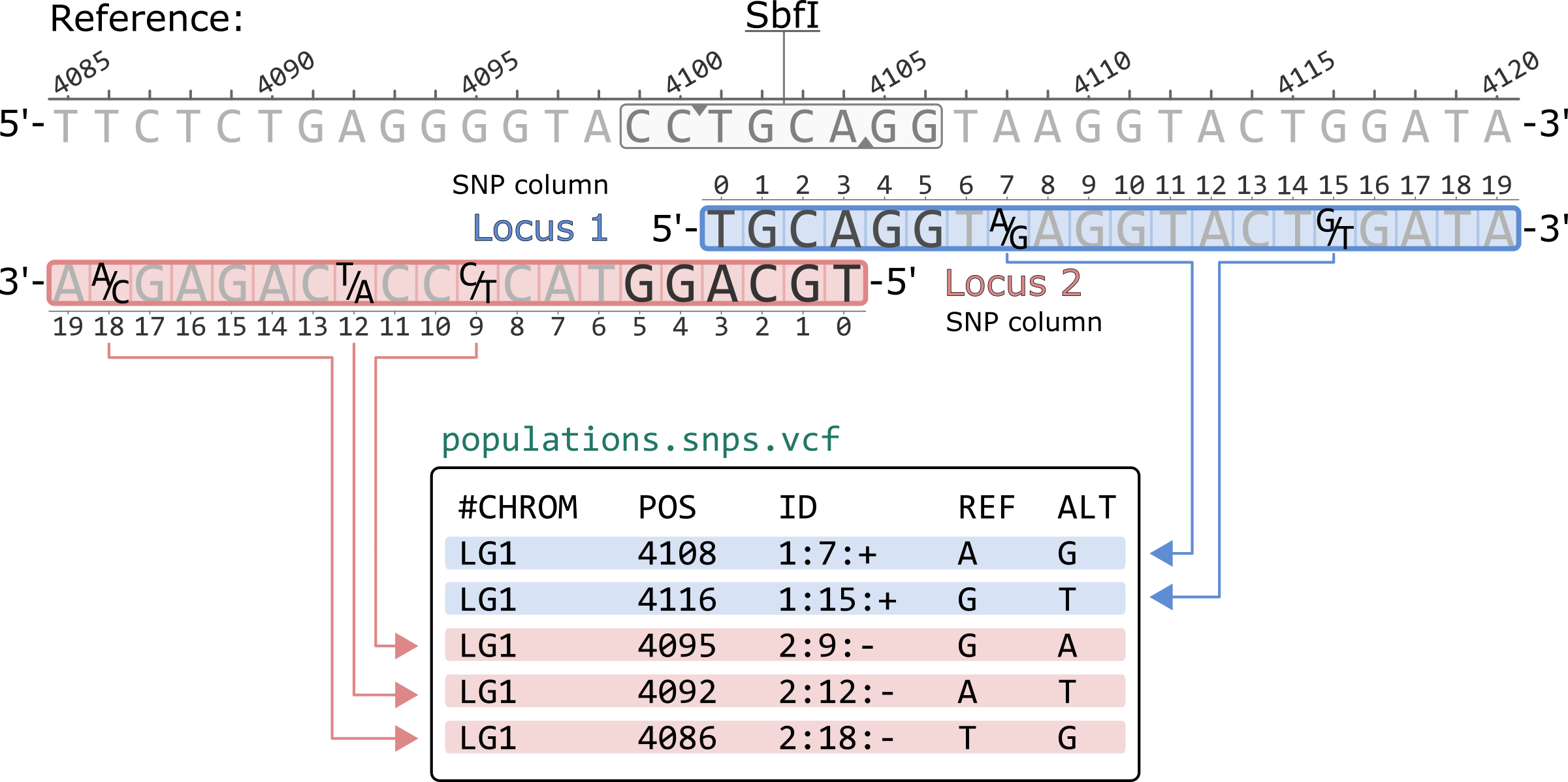
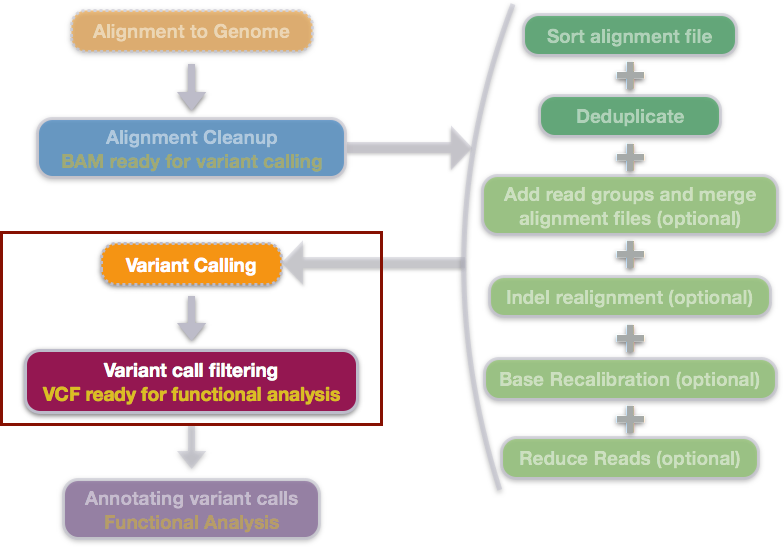
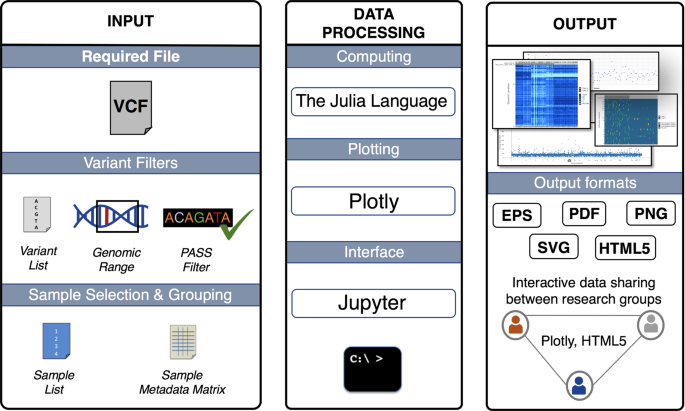
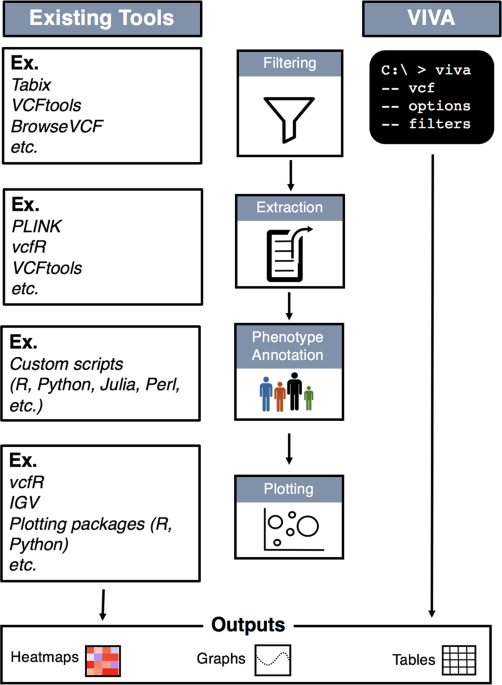
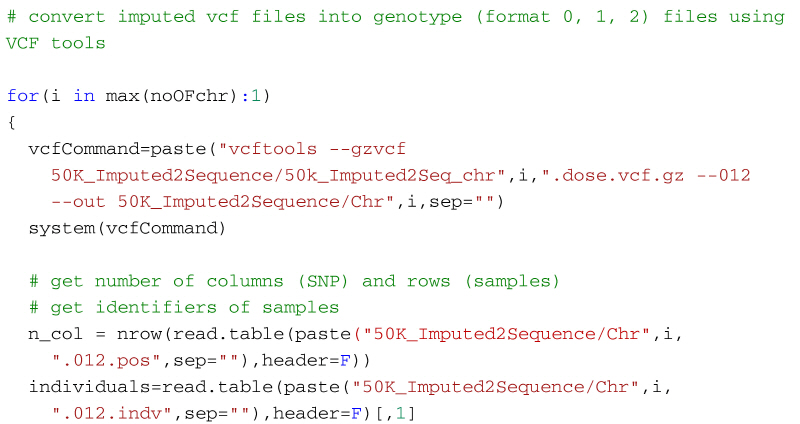
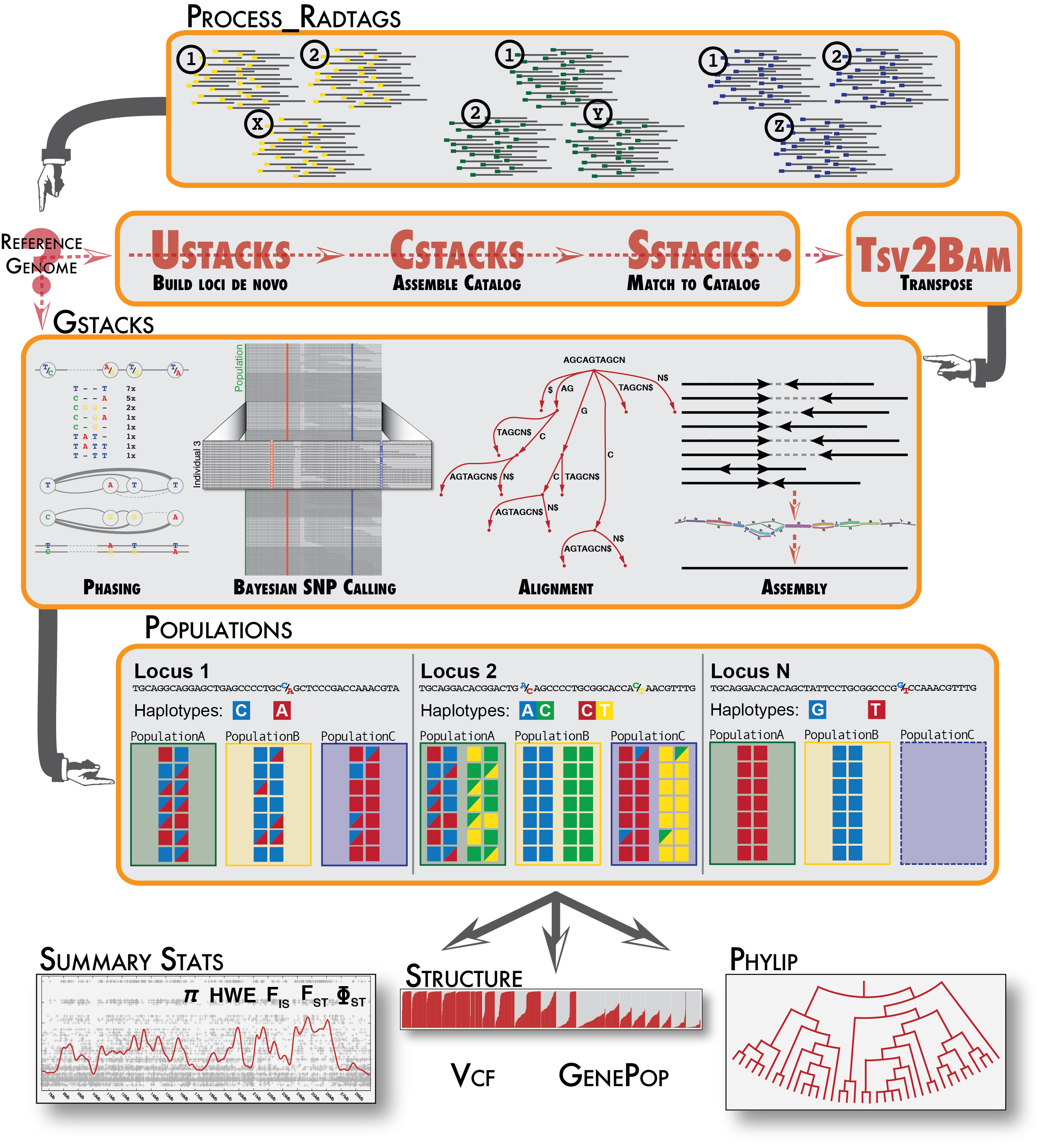
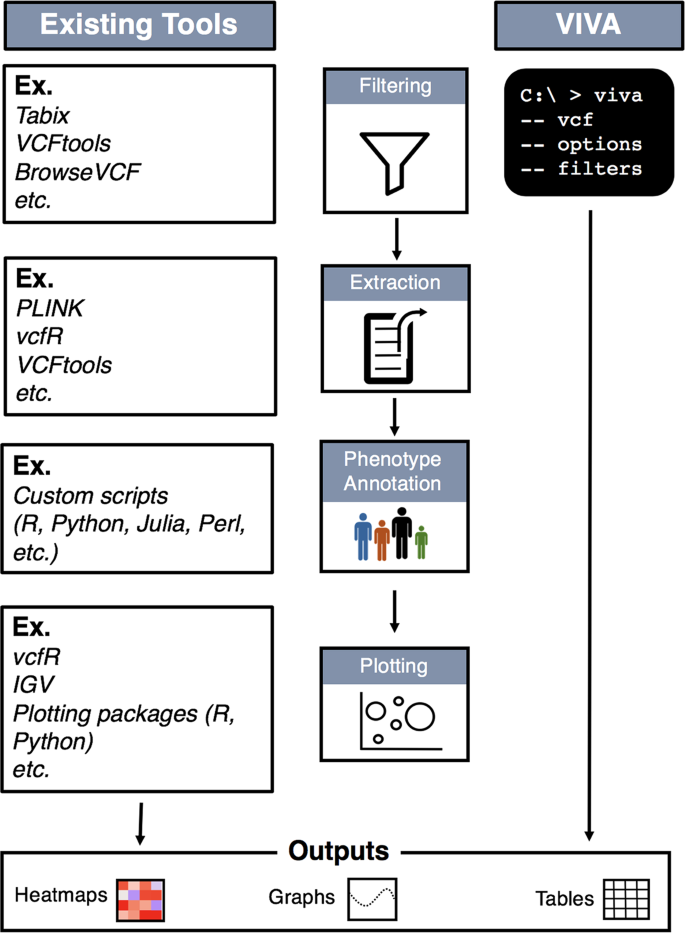
NGS variants output and visualization: VCF and IGV Variant Calling Format (VCF) Variant Calling Format (VCF) Variant Calling For

bcftools view | bcftools tutorial on how to count the number of snps and indels in a vcf file - YouTube
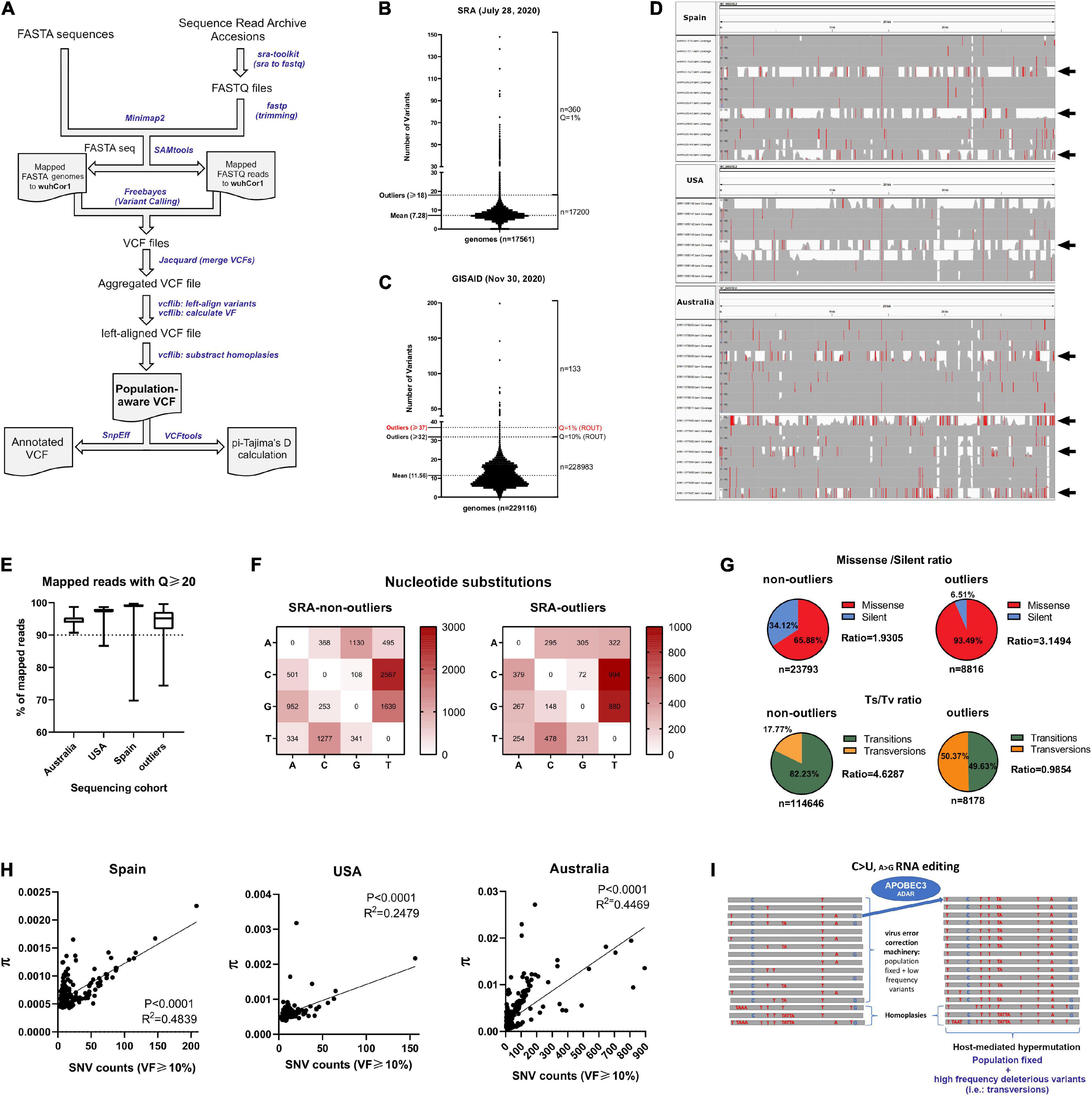
Frontiers | A Novel SARS-CoV-2 Viral Sequence Bioinformatic Pipeline Has Found Genetic Evidence That the Viral 3′ Untranslated Region (UTR) Is Evolving and Generating Increased Viral Diversity