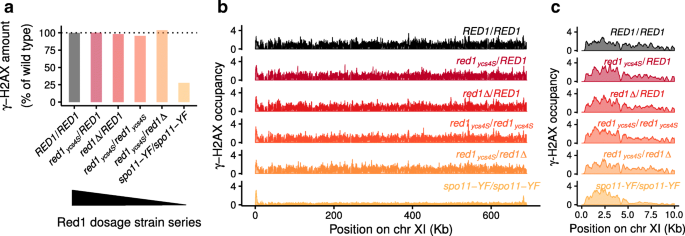
SNP-ChIP: a versatile and tag-free method to quantify changes in protein binding across the genome | BMC Genomics | Full Text
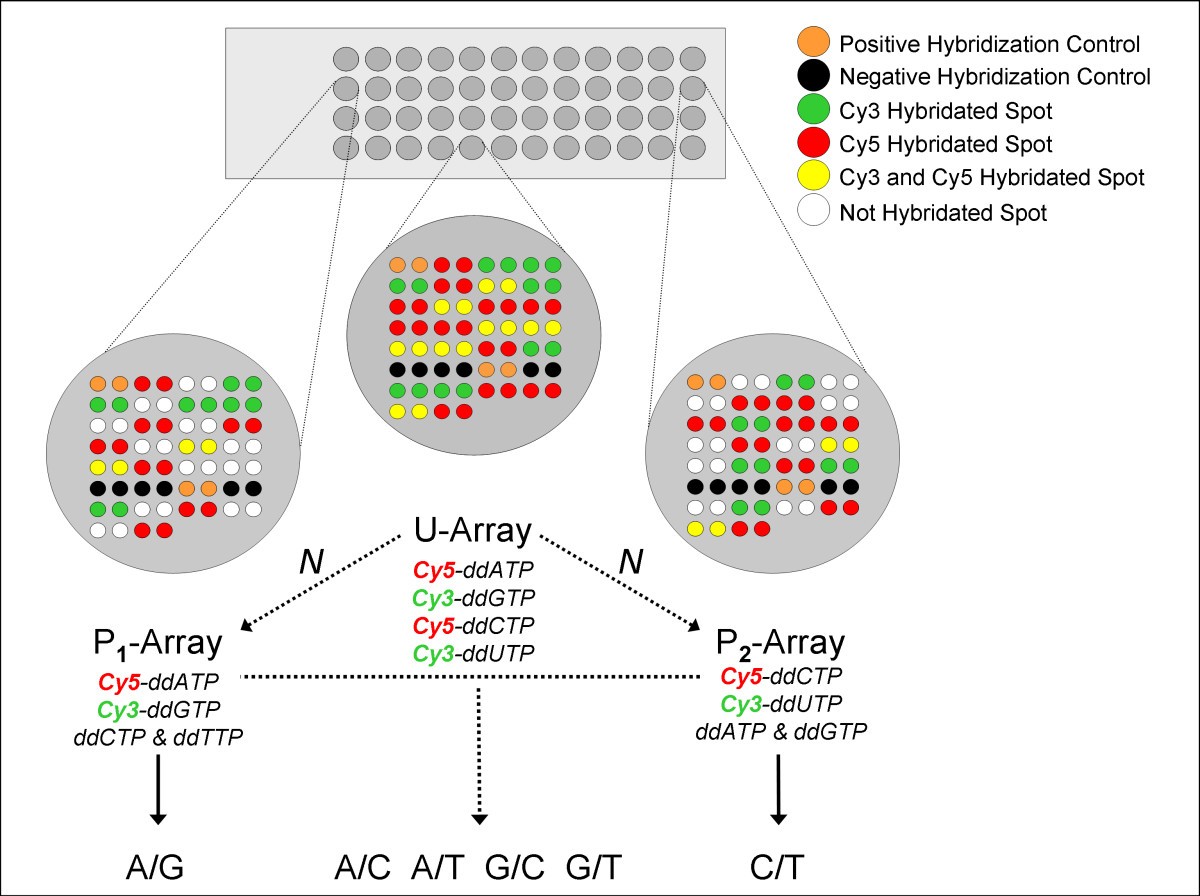
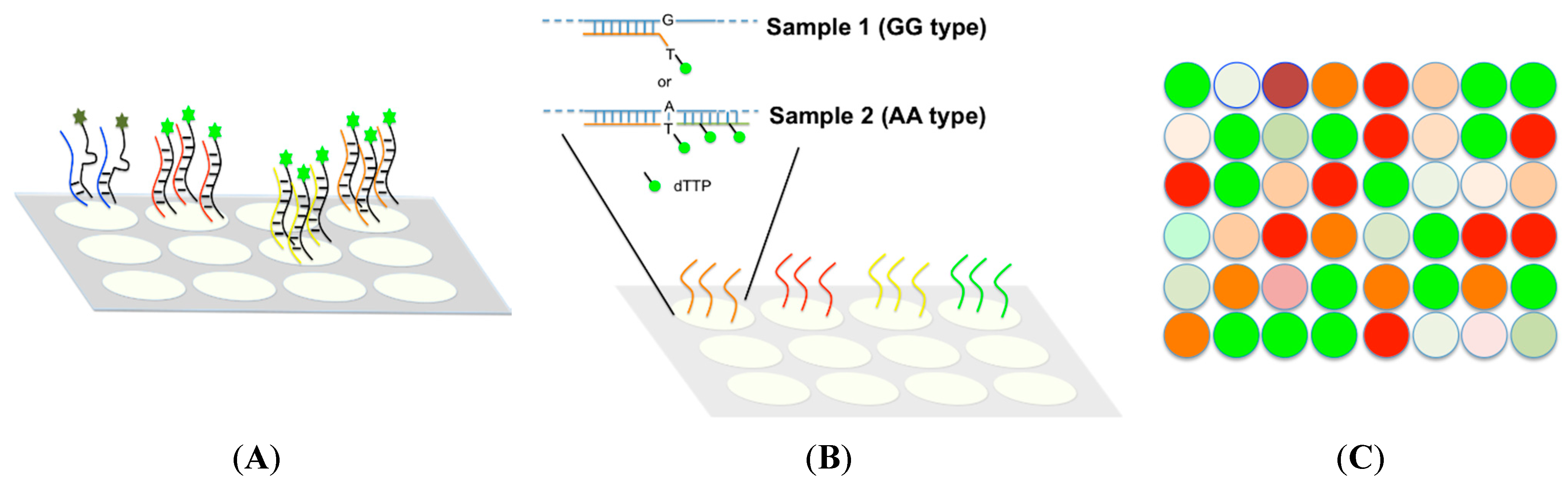
TAMGeS: a Three-Array Method for Genotyping of SNPs by a dual-colour approach | BMC Genomics | Full Text

A typical Lab-on-a-Chip for SNP detection. Biological samples, which... | Download Scientific Diagram

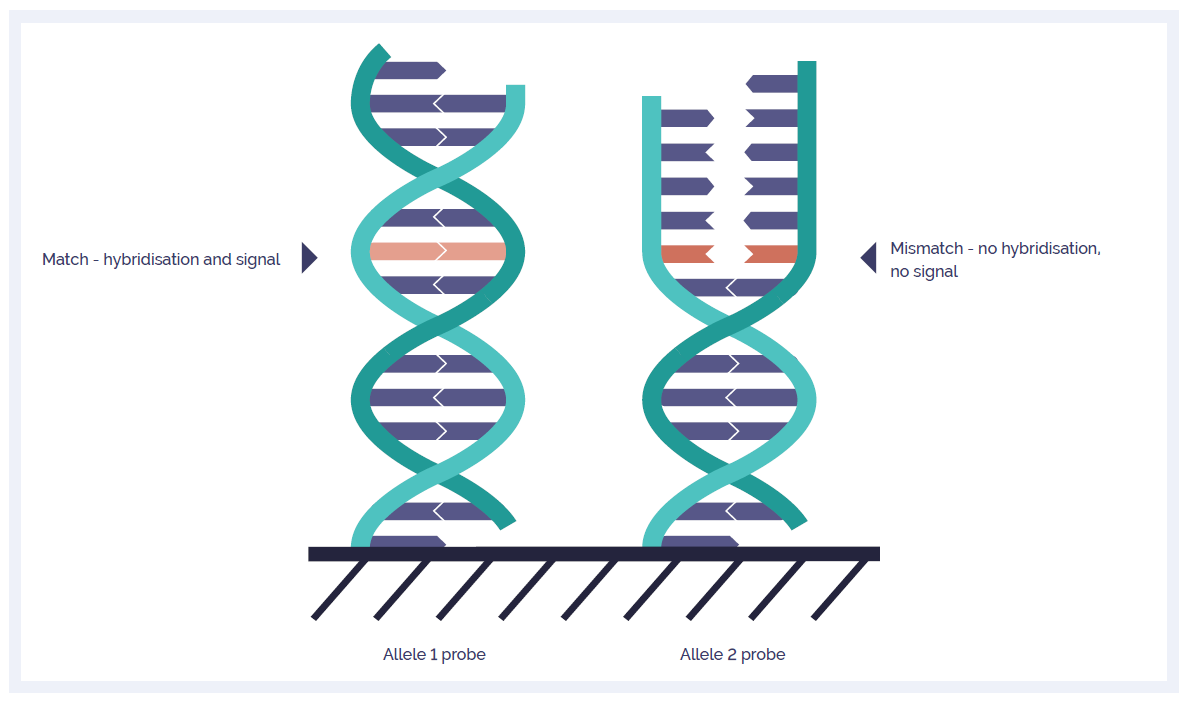
Use of SNP chips to detect rare pathogenic variants: retrospective, population based diagnostic evaluation | The BMJ

A Microfluidic-Based SNP Genotyping Method for Hereditary Hearing-Loss Detection | Analytical Chemistry

3 The GeneChip Genome-Wide Human SNP Array 6.0. The chip is 1.28 by... | Download Scientific Diagram

High-Throughput | Free Full-Text | The Cytoscan HD Array in the Diagnosis of Neurodevelopmental Disorders
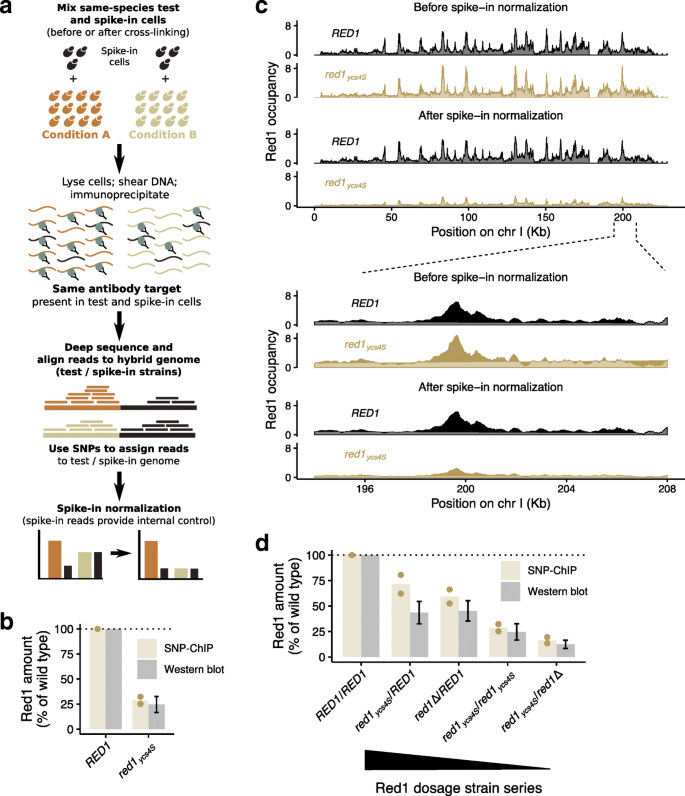
SNP-ChIP: a versatile and tag-free method to quantify changes in protein binding across the genome | BMC Genomics | Full Text

Use of SNP chips to detect rare pathogenic variants: retrospective, population based diagnostic evaluation | The BMJ
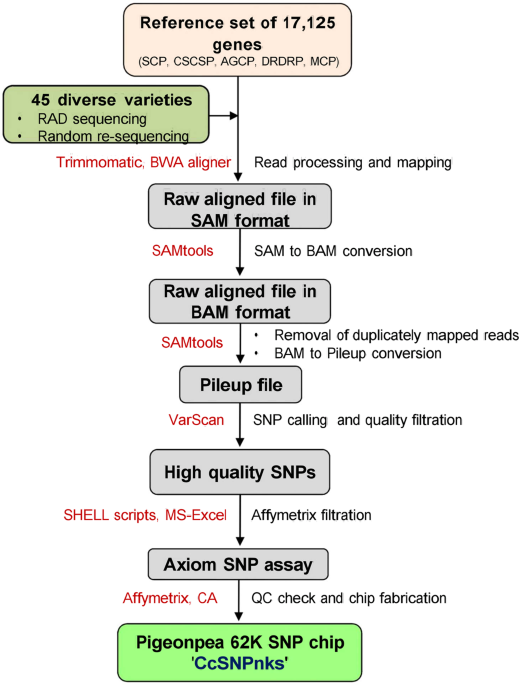
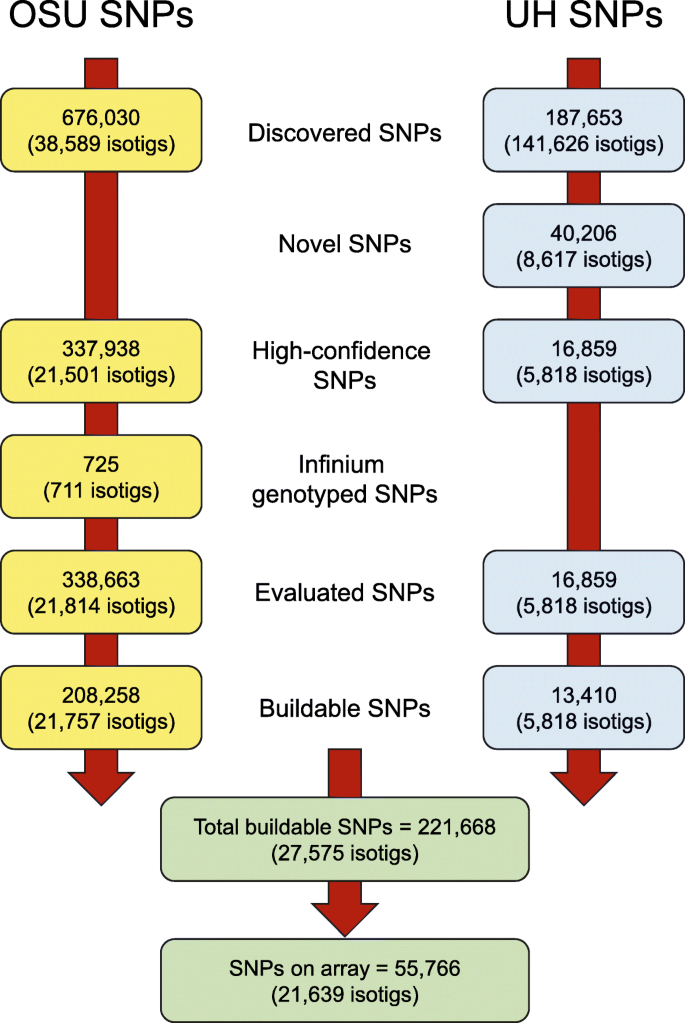
A 62K genic-SNP chip array for genetic studies and breeding applications in pigeonpea (Cajanus cajan L. Millsp.) | Scientific Reports
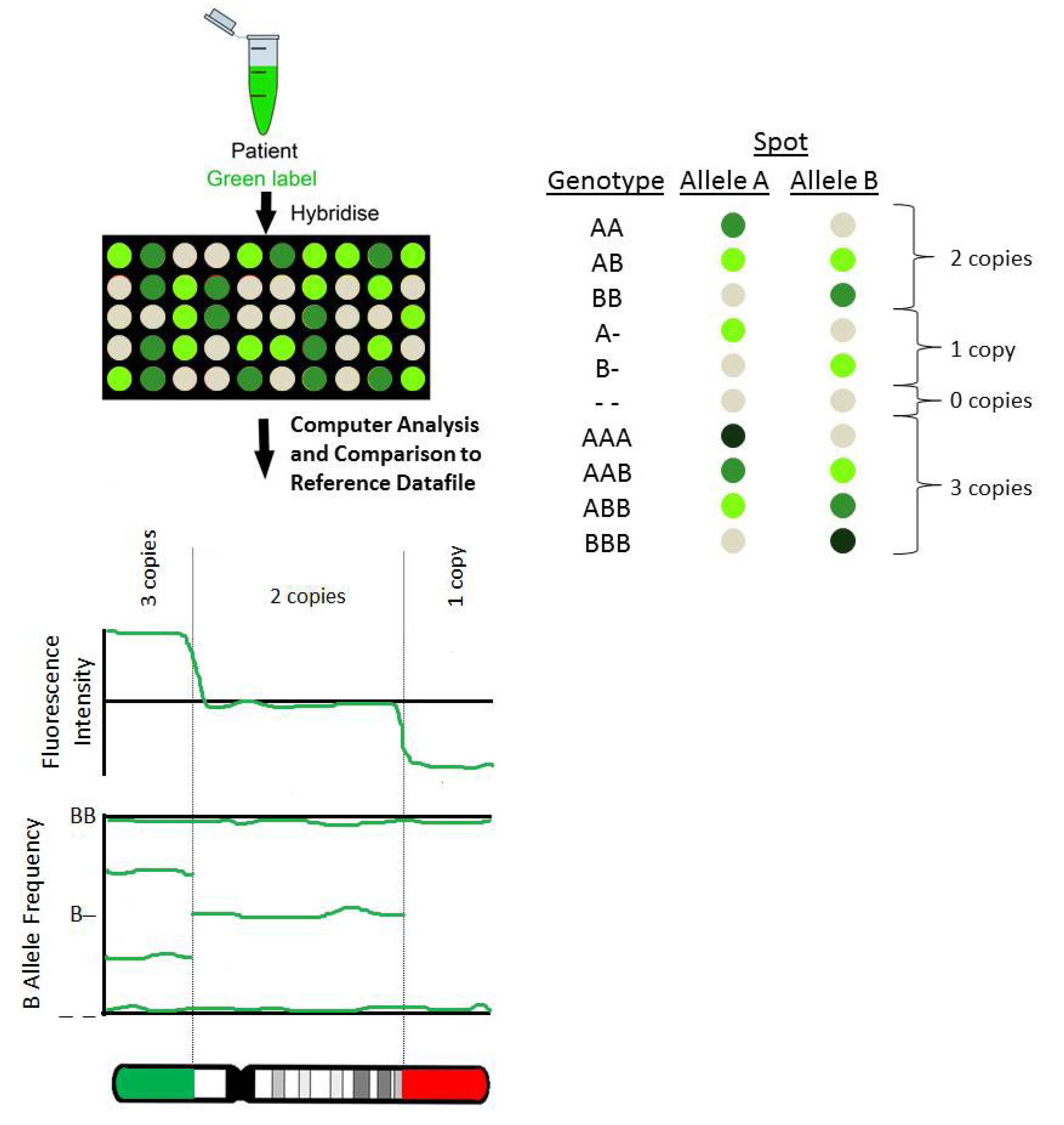
JCM | Free Full-Text | Microarray Technology for the Diagnosis of Fetal Chromosomal Aberrations: Which Platform Should We Use?